|
How many base pairs are in palindromic sequence? It derives from Greek roots that literally mean “running back” (palin is “again, back,” and dromos, “running.”) The word appears to have been created in English based on these roots in the early 1600s. What do you call words that can be read backwards?Ī palindrome is a word, sentence, verse, or even number that reads the same backward or forward. Thus, the name ‘satellite DNA’ was coined. The density of DNA is a function of its base and sequence, and satellite DNA with its highly repetitive DNA has a reduced or a characteristic density compared to the rest of the genome. In the DNA and RNA worlds the term means that one strand reads the same in the 5′ → 3′ direction as the complementary strand reads in the 5′ → 3′ direction. What is a palindrome in DNA and what is one function?Ī palindrome, such as the famous “A man, a plan, a canal, Panama,” reads the same in both directions. Some examples of palindromic words are redivider, deified, civic, radar, level, rotor, kayak, reviver, racecar, madam, and refer. The characters read the same backward as forward. … This is the property that gives CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) its tongue-twisting name. Parts of the letter sequence of one strand (green) correspond to those of the other strand (yellow) in the reverse order. Palindrome sequence in the DNA of the bacterium Streptococcus agalactiae. drome | ˈpal-ən-ˌdrōm What is palindrome number?Ī palindromic number (also known as a numeral palindrome or a numeric palindrome) is a number (such as 16461) that remains the same when its digits are reversed.: a word, phrase, sentence, or number that reads the same backward or forward “Step on no pets” is a palindrome. …Īdvertisements What is the meaning palindromic? For example the recognition sequence for BamHI is GGATCC. These enzymes predictably cut both strands because the sequences they recognize are palindromic. Restriction enzymes cut double-stranded DNA * at specific locations based the pattern of bases found at those locations. What Is a DNA Palindrome? A palindromic sequence of nucleotides (which are labeled A, T, C, or G) occurs when complementary strands of DNA read the same in both directions, either from the 5-prime end or the 3-prime end. Palindromic sequences: nucleic acid sequences where the one strand matches its complementary strand when read in the same direction. 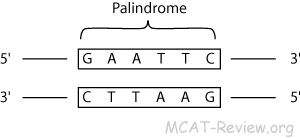
For example, the sequence 5′-CGATCG-3′ is considered a palindrome since its reverse complement 3′-GCTAGC-5′ reads the same. How do you find the palindromic sequence of DNA?įor a nucleotide sequence to be considered as a palindrome, its complementary strand must read the same in the opposite direction. It is also known as a palindrome or an inverted-reverse sequence. What is the structure of palindromic sequence?Ī palindromic sequence is a sequence made up of nucleic acids within double helix of DNA and/or RNA that is the same when read from 5′ to 3′ on one strand and 5′ to 3′ on the other, complementary, strand. In these cases, two different segments of the double helix read the same but in opposite directions. Palindromes that occur on opposite strands of the same section of DNA helix.What are the types of palindromic sequence? Importance of this sequence is that it reads the same in both directions. Palindromic sequences are typically 3 to 5 bases in length. The palindromic sequence has an important role in molecular biology, as the DNA sequence is double-stranded and by reading base pairs palindromes can be determined. What is the importance of palindromic sequence? 5′ to 3′) on one strand matches the sequence reading in the opposite direction (e.g. What is palindromic sequence give examples?Ī palindromic sequence is a nucleic acid sequence in a double-stranded DNA or RNA molecule wherein reading in a certain direction (e.g. It proves to be an unusual enzyme, clearly related functionally to Type II endonuclease. The HsaI restriction enzyme from the embryos of human, Homo sapiens, has been isolated with both the tissue extract and nuclear extract. Today, scientists recognize three categories of restriction enzymes: type I, which recognize specific DNA sequences but make their cut at seemingly random sites that can be as far as 1,000 base pairs away from the recognition site type II, which recognize and cut directly within the recognition site and type III, … Do humans have restriction enzymes? What are the three types of restriction enzymes? These regions are called recognition sequences, or recognition sites, and are randomly distributed throughout the DNA. Each restriction enzyme recognizes a short, specific sequence of nucleotide bases (the four basic chemical subunits of the linear double-stranded DNA molecule-adenine, cytosine, thymine, and guanine).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
